
|
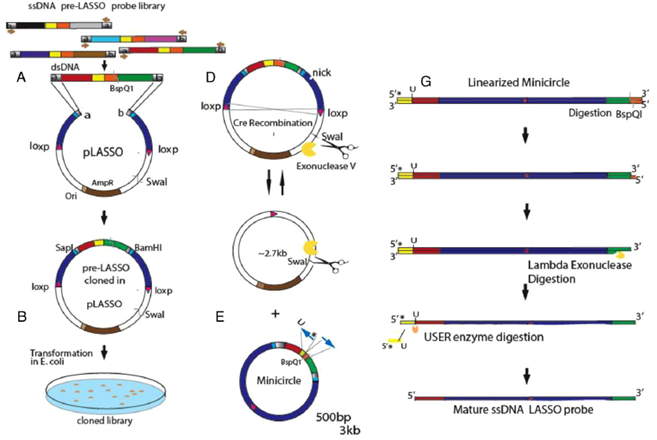
Overall workflow for preparing the LASSO library.
|
Invention Summary:
Advances in DNA sequencing have led to an exponential increase in the quantity of sequence data, whereas limited achievable maximum gene length (<200 bp) can produce highly biased libraries with constraints on sequence homology. Long-adapter single-strand oligonucleotide (LASSO) probes libraries were thus developed to allow highly multiplexed amplification of thousands of kilobase-sized DNA targets in the same reaction volume available for downstream long read DNA sequencing or expression.
Rutgers scientists have developed a rapid, inexpensive method to assemble LASSO probe libraries using a Cre-recombinase-mediated reaction and a protocol for multiplex genome target capture. This strategy generates high quality LASSO probes-based libraries that can targets thousands of DNA regions in a single reaction. A proof-of-concept study has demonstrated that an Escherichia coli open reading frames (ORFs) library generated by this method comprises DNA fragments spanning from 400 bp to 5000 bp. The median enrichment of all targeted ORFs versus untargeted ORFs was 30 times. For ORFs up to 1 kb in size, targeted ORFs were enriched up to a median of 260-fold. Assembly of a LASSO probe library takes 4 days, and target capture can be performed in a single day.
Advantages:
- Rapid, inexpensive
- Captured DNA fragments ranging from few hundred bp to few thousand bp, and potentially much larger fragments
- Robust in DNA fragment coverage
Market Applications:
- Generation of genomic DNA libraries for long-read sequencing
- Multiplexed cloning and expression
Publications:
1. Tosi et.al. Long-adapter single-strand oligonucleotide probes for the massively multiplexed cloning of kilobase genome regions. Nat. Biomed. Eng. 2017; 1:0092
2. Tosi et.al. Massively parallel DNA target capture using long adapter single stranded oligonucleotide (LASSO) probes assembled through a novel DNA recombinase mediated methodology. Biotechnol. J. 2022; 17:2100240.
3. Chkaiban et.al. Assembly of long-adapter single-strand oligonucleotide (LASSO) probes for massively parallel capture of kilobase size DNA targets. Current Protocols, 1, e278, doi: 10.1002/cpz1.278.
Intellectual Property & Development Status: Patent pending. Available for licensing and/or research collaboration.