
Invention Summary:
Determining resistance to chemotherapy treatments for specific types of cancer (e.g., prostate, lung, breast, etc.,) as early as possible can prolong patients' lives by enabling them to be treated with other, and hopefully, more effective treatments as soon as possible.
Researchers at Rutgers University have identified several bioinformatic methods for the identification of biomarkers of resistance to specific treatments for specific types of cancer. These algorithms are:
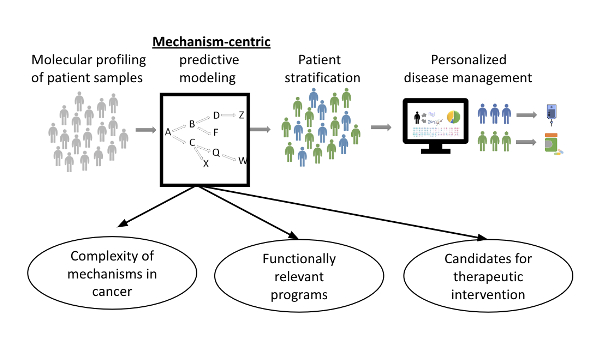
- TR-2-Path – a de novo mechanism-centric regulatory network that encodes relationships between molecular pathways and their upstream transcriptional regulatory programs that have identified markers to predict patients' response to enzalutamide in prostate cancer.
- pathCHEMO: a generalizable genome-wide computational framework that uncovers interplay between transcriptomic and epigenomic mechanisms altered in biological pathways that has been successfully used to identify seven molecular pathway biomarkers for resistance to adjuvant standard-of-care doublet chemotherapy (i.e., carboplatin-paclitaxel) in lung adenocarcinoma.
- Epi2Genr: a systematic genome-wide computational method which integrates DNA methylation and mRNA expression data that successfully uncovered a panel of 5 differentially methylated sites able to accurately predict response to ADT across independent prostate cancer cohorts.
Advantages:
- Algorithms facilitate the identification of new biomarkers of resistance for cancers.
- Improve clinical decision-making, avoid harmful side effects, patients’ survival, and cancer management.
- Strong foundation for personalized medicine
Market Applications:
- Identification of new biomarkers for drug resistance for specific cancer types
- Use of biomarkers to determine/recommend chemotherapeutic treatments for specific cancers.
Intellectual Property & Development Status: Provisional patent application filed. Available for licensing and/or search collaboration. For any business development and other collaborative partnerships contact marketingbd@research.rutgers.edu